|
[1]
|
 Cameron C. Gray, Shatha F. Al-Maliki, & Franck P. Vidal.
Data exploration in evolutionary reconstruction of PET images.
Genetic Programming and Evolvable Machines, 19(3):391-419,
September 2018.
Cameron C. Gray, Shatha F. Al-Maliki, & Franck P. Vidal.
Data exploration in evolutionary reconstruction of PET images.
Genetic Programming and Evolvable Machines, 19(3):391-419,
September 2018.



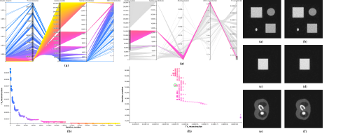
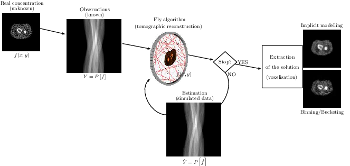
This work is based on a cooperative co-evolution algorithm called
`Fly Algorithm', which is an evolutionary algorithm (EA) where individuals
are called `flies'. It is a specific case of the `Parisian Approach' where
the solution of an optimisation problem is a set of individuals
(e.g. the whole population) instead of a single individual (the best one) as
in typical EAs. The optimisation problem considered here is tomography
reconstruction in positron emission tomography (PET). It estimates
the concentration of a radioactive substance (called a radiotracer) within
the body. Tomography, in this context, is considered as a difficult
ill-posed inverse problem. The Fly Algorithm aims at optimising
the position of 3-D points that mimic the radiotracer. At the end of
the optimisation process, the fly population is extracted as it corresponds
to an estimate of the radioactive concentration. During the optimisation
loop a lot of data is generated by the algorithm, such as image metrics,
duration, and internal states. This data is recorded in a log file that
can be post-processed and visualised. We propose using information
visualisation and user interaction techniques to explore the algorithm's
internal data. Our aim is to better understand what happens during
the evolutionary loop. Using an example, we demonstrate that it is possible
to interactively discover when an early termination could be triggered.
It is implemented in a new stopping criterion. It is tested on two other
examples on which it leads to a 60% reduction of the number of iterations
without any loss of accuracy.
Keywords: Fly Algorithm; Tomography reconstruction; Information visualisation; Data exploration; Artificial evolution; Parisian evolution.
|
|
[2]
|
 Z. Ali Abbood, J. Lavauzelle, É Lutton, J.-M. Rocchisani, J. Louchet, and F. P. Vidal.
Voxelisation in the 3-D Fly Algorithm for PET.
Swarm and Evolutionary Computation, 36:91-105,
October 2017.
Z. Ali Abbood, J. Lavauzelle, É Lutton, J.-M. Rocchisani, J. Louchet, and F. P. Vidal.
Voxelisation in the 3-D Fly Algorithm for PET.
Swarm and Evolutionary Computation, 36:91-105,
October 2017.



The Fly Algorithm was initially developed for 3-D robot vision applications. It consists in solving the inverse problem of
shape reconstruction from projections by evolving a population of 3-D points in space (the `flies'), using an
evolutionary optimisation strategy. Here, in its version dedicated to tomographic reconstruction in medical imaging, the
flies are mimicking radioactive photon sources. Evolution is controlled using a fitness function based on the
discrepancy of the projections simulated by the flies with the actual pattern received by the sensors. The reconstructed
radioactive concentration is derived from the population of flies, i.e. a collection of points in the 3-D Euclidean space,
after convergence. `Good' flies were previously binned into voxels. In this paper, we study which flies to include in the
final solution and how this information can be sampled to provide more accurate datasets in a reduced computation
time. We investigate the use of density fields, based on Metaballs and on Gaussian functions respectively, to obtain a
realistic output. The spread of each Gaussian kernel is modulated in function of the corresponding fly fitness. The
resulting volumes are compared with previous work in terms of normalised-cross correlation. In our test-cases, data
fidelity increases by more than 10% when density fields are used instead of binning. Our method also provides
reconstructions comparable to those obtained using well-established techniques used in medicine (filtered back-
projection and ordered subset expectation-maximisation).
Keywords: Fly algorithm; Evolutionary computation; tomography reconstruction; iterative
algorithms; inverse problems; co-operative co-evolution
|
|
[3]
|
 F. P. Vidal, P.-F. Villard, and É. Lutton.
Tuning of patient specific deformable models using an adaptive
evolutionary optimization strategy.
IEEE Transactions on Biomedical Engineering, 59(10):2942-2949,
October 2012.
F. P. Vidal, P.-F. Villard, and É. Lutton.
Tuning of patient specific deformable models using an adaptive
evolutionary optimization strategy.
IEEE Transactions on Biomedical Engineering, 59(10):2942-2949,
October 2012.




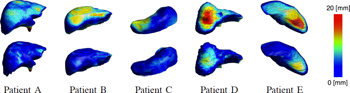
We present and analyze the behavior of an evolutionary algorithm designed to estimate the parameters of a complex organ behavior model. The model is adaptable to account for patient's specificities. The aim is to finely tune the model to be accurately adapted to various real patient datasets. It can then be embedded, for example, in high fidelity simulations of the human physiology. We present here an application focused on respiration modeling. The algorithm is automatic and adaptive. A compound fitness function has been designed to take into account for various quantities that have to be minimized. The algorithm efficiency is experimentally analyzed on several real test cases: 1) three patient datasets have been acquired with the "breath hold" protocol, and 2) two datasets corresponds to 4-D CT scans. Its performance is compared with two traditional methods (downhill simplex and conjugate gradient descent): a random search and a basic real-valued genetic algorithm. The results show that our evolutionary scheme provides more significantly stable and accurate results.
Keywords: Evolutionary computation, inverse problems, medical simulation,
adaptive algorithm
|
|
[1]
|
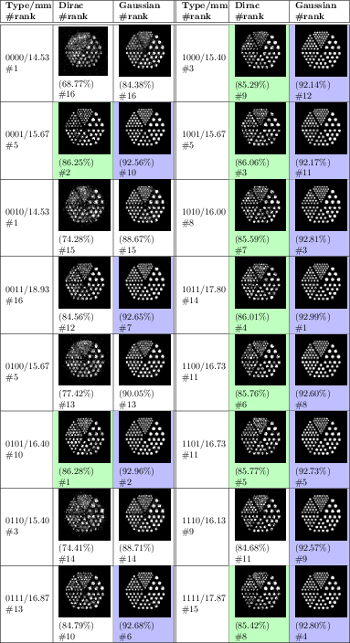 Zainab Ali Abbood, and Franck P. Vidal. Basic, dual, adaptive, and directed mutation operators in the Fly algorithm.
In Biennial International Conference on Artificial Evolution, volume ???? of
Lecture Notes in Computer Science, pages ???-???, Paris, France,
October 2017. Springer, Heidelberg.
Zainab Ali Abbood, and Franck P. Vidal. Basic, dual, adaptive, and directed mutation operators in the Fly algorithm.
In Biennial International Conference on Artificial Evolution, volume ???? of
Lecture Notes in Computer Science, pages ???-???, Paris, France,
October 2017. Springer, Heidelberg.



Our work is based on a Cooperative Co-evolution Algorithm -- the Fly algorithm -- in which individuals correspond to 3-D points. The Fly algorithm uses two levels of fitness function: i) a local fitness computed to evaluate a given individual (usually during the selection process) and ii) a global fitness to assess the performance of the population as a whole. This global fitness is the metrics that is minimised (or maximised depending on the problem) by the optimiser. Here the solution of the optimisation problem corresponds to a set of individuals instead of a single individual (the best individual) as in classical evolutionary algorithms. The Fly algorithm heavily relies on mutation operators and a new blood operator to insure diversity in the population. To lead to accurate results, a large mutation variance is often initially used to avoid local minima (or maxima). It is then progressively reduced to refine the results. Another approach is the use of adaptive operators. However, very little research on adaptive operators in Fly algorithm has been conducted. We address this deficiency and propose 4 different fully adaptive mutation operators in the Fly algorithm: positrons, and the final solution of the algorithm approximates the radioactivity concentration. The view and analysis four mutation operators, which are Basic Mutation, Adaptive Mutation Variance, Dual Mutation, and Directed Mutation. Due to the complex nature of the search space, (kN-dimensions, with k the number of genes per individuals and N the number of individuals in the population), we favour operators with a low maintenance cost in terms of computations. Their impact on the algorithm efficiency is analysed and validated on positron emission tomography (PET) reconstruction.
Keywords: evolutionary algorithms, Parisian approach, reconstruction algorithms, positron emission tomography, mutation operator
|
|
[2]
|
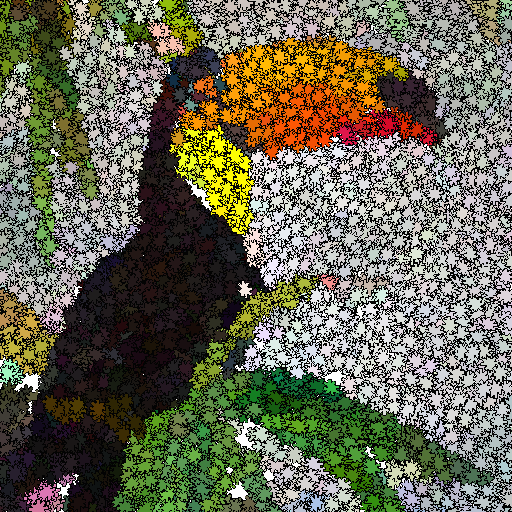 Z. Ali Abbood, O. Amlal, and F. P. Vidal.
Evolutionary Art Using the Fly Algorithm.
In Applications of Evolutionary Computation, volume 10199 of
Lecture Notes in Computer Science, pages 455-470, Amsterdam, The Netherlands,
April 2017. Springer, Heidelberg.
Z. Ali Abbood, O. Amlal, and F. P. Vidal.
Evolutionary Art Using the Fly Algorithm.
In Applications of Evolutionary Computation, volume 10199 of
Lecture Notes in Computer Science, pages 455-470, Amsterdam, The Netherlands,
April 2017. Springer, Heidelberg.



This study is about Evolutionary art such as digital mosaics. The most common techniques to generate a digital mosaic effect heavily rely on Centroidal Voronoi diagrams. Our method generates artistic images as an optimisation problem without the introduction of any a priori knowledge or constraint other than the input image. We adapt a cooperative co-evolution strategy based on the Parisian evolution approach, the Fly algorithm, to produce artistic visual effects from an input image (e.g. a photograph). The primary usage of the Fly algorithm is in computer vision, especially stereo-vision in robotics. It has also been used in image reconstruction for tomography. Until now the individuals correspond to simplistic primitives: Infinitely small 3-D points. In this paper, the individuals have a much more complex representation and represent tiles in a mosaic. They have their own position, size, colour, and rotation angle. We take advantage of graphics processing units (GPUs) to generate the images using the modern OpenGL Shading Language. Different types of tiles are implemented, some with transparency, to generate different visual effects, such as digital mosaic and spray paint. A user study has been conducted to evaluate some of our results. We also compare results with those obtained with GIMP, an open-source software for image manipulation.
Keywords: Digital mosaic; Evolutionary art; Fly algorithm; Parisian evolution; Cooperative co-evolution
|
|
[3]
|
 F. P. Vidal, Y. L. Pavia, J.-M. Rocchisani, J. Louchet, and É. Lutton.
Artificial evolution strategy for pet reconstruction.
In International Conference on Medical Imaging Using
Bio-Inspired and Soft Computing (MIBISOC2013), pages 39-46, Brussels,
Belgium, May 2013.
F. P. Vidal, Y. L. Pavia, J.-M. Rocchisani, J. Louchet, and É. Lutton.
Artificial evolution strategy for pet reconstruction.
In International Conference on Medical Imaging Using
Bio-Inspired and Soft Computing (MIBISOC2013), pages 39-46, Brussels,
Belgium, May 2013.


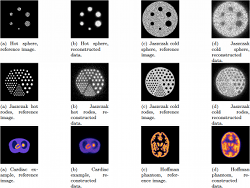
This paper shows new resutls of our artificial evolution algorithm
for Positron Emission Tomography (PET) reconstruction. This imaging technique
produces datasets corresponding to the concentration of positron emitters
within the patient. Fully three-dimensional (3D) tomographic reconstruction
requires high computing power and leads to many challenges. Our aim is
to produce high quality datasets in a time that is clinically acceptable.
Our method is based on a co-evolution strategy called the “Fly algorithm”.
Each fly represents a point in space and mimics a positron emitter. Each fly
position is progressively optimised using evolutionary computing to closely
match the data measured by the imaging system. The performance of
each fly is assessed based on its positive or negative contribution to
the performance of the whole population. The final population of flies
approximates the radioactivity concentration. This approach has shown
promising results on numerical phantom models. The size of objects and
their relative concentrations can be calculated in two-dimensional (2D)
space. In (3D), complex shapes can be reconstructed. In this paper,
we demonstrate the ability of the algorithm to fidely reconstruct more
anatomically realistic volumes.
Keywords: Evolutionary computation, inverse problems, adaptive algorithm, Nuclear medicine, Positron emission tomography, Reconstruction algorithms
|
|
[4]
|
 F. P. Vidal, P.-F. Villard, and É. Lutton.
Automatic tuning of respiratory model for patient-based simulation.
In International Conference on Medical Imaging Using
Bio-Inspired and Soft Computing (MIBISOC2013), pages 225-231, Brussels,
Belgium, May 2013.
F. P. Vidal, P.-F. Villard, and É. Lutton.
Automatic tuning of respiratory model for patient-based simulation.
In International Conference on Medical Imaging Using
Bio-Inspired and Soft Computing (MIBISOC2013), pages 225-231, Brussels,
Belgium, May 2013.


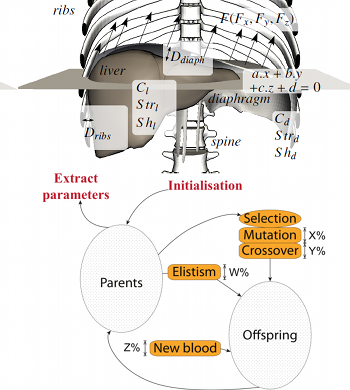
This paper is an overview of a method recently published
in a biomedical journal (IEEE Transactions on Biomedical
Engineering, http://tbme.embs.org). The method is based
on an optimisation technique
called “evolutionary strategy” and it has been designed to
estimate the parameters of a complex 15-D respiration model.
This model is adaptable to account for patient's specificities.
The aim of the optimisation algorithm is to finely tune the
model so that it accurately fits real patient datasets. The final
results can then be embedded, for example, in high fidelity
simulations of the human physiology. Our algorithm is fully
automatic and adaptive. A compound fitness function has been
designed to take into account for various quantities that have
to be minimised (here topological errors of the liver and the
diaphragm geometries). The performance our implementation is
compared with two traditional methods (downhill simplex and
conjugate gradient descent), a random search and a basic real-valued
genetic algorithm. It shows that our evolutionary scheme
provides results that are significantly more stable and accurate
than the other tested methods. The approach is relatively generic
and can be easily adapted to other complex parametrisation
problems when ground truth data is available.
Keywords: Evolutionary computation, inverse problems, medical simulation, adaptive algorithm
|
|
[5]
|
 P.-F. Villard, F. P. Vidal, F. Bello, and N. W. John.
A method to compute respiration parameters for patient-based
simulators.
In Proceeding of Medicine Meets Virtual Reality 19 - NextMed
(MMVR19), volume 173 of Studies in Health Technology and Informatics,
pages 529-533, Newport Beach, California, February 2012. IOS Press.
Winner of the best poster award.
P.-F. Villard, F. P. Vidal, F. Bello, and N. W. John.
A method to compute respiration parameters for patient-based
simulators.
In Proceeding of Medicine Meets Virtual Reality 19 - NextMed
(MMVR19), volume 173 of Studies in Health Technology and Informatics,
pages 529-533, Newport Beach, California, February 2012. IOS Press.
Winner of the best poster award.



We propose a method to automatically tune a patient-based virtual environment training
simulator for abdominal needle insertion. The key attributes to be customized in our framework are
the elasticity of soft-tissues and the respiratory model parameters. The estimation is based on two
3D Computed Tomography (CT) scans of the same patient at two different time steps. Results are
presented on five patients and show that our new method leads to better results than our previous
studies with manually tuned parameters.
|
|
[6]
|
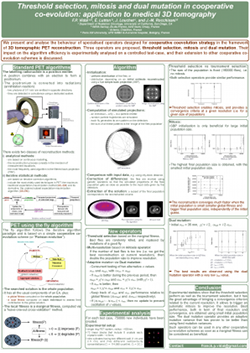 F. P. Vidal, É. Lutton, J. Louchet, and J.-M. Rocchisani.
Threshold selection, mitosis and dual mutation in cooperative
coevolution: application to medical 3D tomography.
In International Conference on Parallel Problem Solving From
Nature (PPSN'10), volume 6238 of Lecture Notes in Computer Science,
pages 414-423, Krakow, Poland, September 2010. Springer, Heidelberg.
F. P. Vidal, É. Lutton, J. Louchet, and J.-M. Rocchisani.
Threshold selection, mitosis and dual mutation in cooperative
coevolution: application to medical 3D tomography.
In International Conference on Parallel Problem Solving From
Nature (PPSN'10), volume 6238 of Lecture Notes in Computer Science,
pages 414-423, Krakow, Poland, September 2010. Springer, Heidelberg.



We present and analyse the behaviour of specialised operators
designed for cooperative coevolution strategy in the framework of
3D tomographic PET reconstruction. The basis is a simple cooperative
co-evolution scheme (the “fly algorithm”), which embeds the searched
solution in the whole population, letting each individual be only a part
of the solution. An individual, or fly, is a 3D point that emits positrons.
Using a cooperative co-evolution scheme to optimize the position of
positrons, the population of flies evolves so that the data estimated from
flies matches measured data. The final population approximates the radioactivity
concentration. In this paper, three operators are proposed,
threshold selection, mitosis and dual mutation, and their impact on the
algorithm efficiency is experimentally analysed on a controlled test-case.
Their extension to other cooperative co-evolution schemes is discussed.
|
|
[7]
|
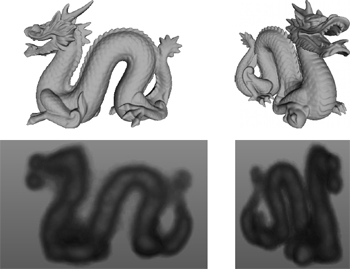 F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
New genetic operators in the Fly algorithm: application to medical
PET image reconstruction.
In Applications of Evolutionary Computation, volume 6024 of
Lecture Notes in Computer Science, pages 292-301, Istanbul, Turkey,
April 2010. Springer, Heidelberg.
Nominated for best paper award.
F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
New genetic operators in the Fly algorithm: application to medical
PET image reconstruction.
In Applications of Evolutionary Computation, volume 6024 of
Lecture Notes in Computer Science, pages 292-301, Istanbul, Turkey,
April 2010. Springer, Heidelberg.
Nominated for best paper award.



This paper presents an evolutionary approach for image reconstruction
in positron emission tomography (PET). Our reconstruction
method is based on a cooperative coevolution strategy (also called
Parisian evolution): the “fly algorithm”. Each fly is a 3D point that
mimics a positron emitter. The flies' position is progressively optimised
using evolutionary computing to closely match the data measured by
the imaging system. The performance of each fly is assessed using a
“marginal evaluation” based on the positive or negative contribution of
this fly to the performance of the population. Using this property, we
propose a “thresholded-selection” method to replace the classical tournament
method. A mitosis operator is also proposed. It is triggered to
automatically increase the population size when the number of flies with
negative fitness becomes too low.
|
|
[8]
|
 F. P. Vidal, D. Lazaro-Ponthus, S. Legoupil, J. Louchet, É. Lutton, and
J.-M. Rocchisani.
Artificial evolution for 3D PET reconstruction.
In Proceedings of the 9th international conference on Artificial
Evolution (EA'09), volume 5975 of Lecture Notes in Computer Science,
pages 37-48, Strasbourg, France, October 2009. Springer, Heidelberg.
F. P. Vidal, D. Lazaro-Ponthus, S. Legoupil, J. Louchet, É. Lutton, and
J.-M. Rocchisani.
Artificial evolution for 3D PET reconstruction.
In Proceedings of the 9th international conference on Artificial
Evolution (EA'09), volume 5975 of Lecture Notes in Computer Science,
pages 37-48, Strasbourg, France, October 2009. Springer, Heidelberg.



This paper presents a method to take advantage of artificial
evolution in positron emission tomography reconstruction. This imaging
technique produces datasets that correspond to the concentration of
positron emitters through the patient. Fully 3D tomographic reconstruction
requires high computing power and leads to many challenges. Our
aim is to reduce the computing cost and produce datasets while retaining
the required quality. Our method is based on a coevolution strategy (also
called Parisian evolution) named “Fly algorithm”. Each fly represents a
point of the space and acts as a positron emitter. The final population of
flies corresponds to the reconstructed data. Using “marginal evaluation”,
the fly's fitness is the positive or negative contribution of this fly to the
performance of the population. This is also used to skip the relatively
costly step of selection and simplify the evolutionary algorithm.
|
|
[1]
|
 F. P. Vidal, M. Folkerts, N. Freud, and S. Jiang.
GPU accelerated DRR computation with scatter.
Medical Physics, 38(6):3455-3456, July 2011.
F. P. Vidal, M. Folkerts, N. Freud, and S. Jiang.
GPU accelerated DRR computation with scatter.
Medical Physics, 38(6):3455-3456, July 2011.



Purpose: We propose a fast software library implemented on
graphics processing unit (GPU) to compute digitally reconstructed
radiographs (DRRs). It takes into account first order Compton
scattering.
Methods: The simulation is based on the evaluation of
the Beer‐Lambert law and of the Klein‐Nishina equation. The
algorithm is fully determinist and has been fully implemented on
GPU to achieve clinically acceptable efficiency. A full resolution
simulation is performed for primary radiation. A much lower image
resolution is used for Compton scattering as it adds a low frequency
pattern over the projection image. Each voxel of the CT dataset is
considered as a secondary source. The number of photons that reach
each voxel is evaluated. Then, for each secondary source, a projection
image is computed and integrated in the final image. The photon energy
between each secondary source and each pixel is also computed.
An interlaced sampling mode is also proposed to further reduce
the computation time without sacrificing numerical accuracy. Finally,
the speed and accuracy are assessed.
Results: We show that the computations can be fully implemented on
the GPU with an original under‐sampling method to produce clinically
acceptable results. For example, a simulation can be achieved in less
than 7 seconds whilst the maximum relative error remains below 5% and
the average relative error below 1.4%. At full resolution, a speed‐up
by factor ∼12X is achieved for the GPU implementation with our
interlaced‐mode by comparison with our multi‐threaded CPU implementation
using 8 threads in parallel.
Conclusions: DRR calculation with scatter is
computationally intensive. The use of GPU can achieve clinically
acceptable efficiency. A Compton fluence map can be computed in a few
seconds using under‐sampling, whilst keeping numerical inaccuracies
relatively low. This work can be used for CBCT reconstruction to reduce
scatter artifacts.
|
|
[2]
|
 F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
Flies for PET: An artificial evolution strategy for image
reconstruction in nuclear medicine.
Medical Physics, 37(6):3139, July 2010.
F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
Flies for PET: An artificial evolution strategy for image
reconstruction in nuclear medicine.
Medical Physics, 37(6):3139, July 2010.



Purpose: We propose an evolutionary approach for image
reconstruction in nuclear medicine. Our method is based on
a cooperative coevolution strategy (also called Parisian evolution):
the “fly algorithm”.
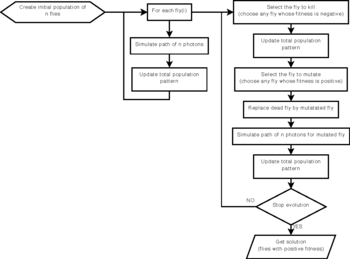
Method and Materials: Each individual, or fly,
corresponds to a 3D point that mimics a radioactive emitter, i.e.
a stochastic simulation of annihilation events is performed to compute
the fly's illumination pattern. For each annihilation, a photon is
emitted in a random direction, and a second photon is emitted in
the opposite direction. The line between two detected photons is
called line of response (LOR). If both photons are detected by
the scanner, the fly's illumination pattern is updated.
The LORs of every fly are aggregated to form the population total
illumination pattern. Using genetic operations to optimize the position
of positrons, the population of flies evolves so that the population
total pattern matches measured data. The final population of flies
approximates the radioactivity concentration.
Results: We have developed numerical phantom models to assess
the reconstruction algorithm. To date, no scattering and no tissue attenuation
have been considered. Whilst this is not physically correct, it allows us
to test and validate our approach in the simplest cases.
Preliminary results show the validity of this approach in both 2D and
fully‐3D modes. In particular, the size of objects, and
their relative concentrations can be retrieved in the 2D mode.
In fully‐3D, complex shapes can be reconstructed.
Conclusions: An evolutionary approach for PET reconstruction has been proposed
and validated using simple test cases. Further work will therefore include
the use of more realistic input data (including random events and scattering),
which will finally lead to implement the correction of scattering within our algorithm.
A comparison study against ML‐EM and/or OS‐EM methods will also need to be conducted.
|
|
[1]
|
 Zainab Ali Abbood, Jean-Marie Rocchisani, and Franck P. Vidal.
Visualisation of PET data in the Fly Algorithm.
In Katja Bühler, Lars Linsen, and Nigel W. John, editors,
Eurographics Workshop on Visual Computing for Biology and Medicine, pages
211--212. The Eurographics Association, 2015.
Zainab Ali Abbood, Jean-Marie Rocchisani, and Franck P. Vidal.
Visualisation of PET data in the Fly Algorithm.
In Katja Bühler, Lars Linsen, and Nigel W. John, editors,
Eurographics Workshop on Visual Computing for Biology and Medicine, pages
211--212. The Eurographics Association, 2015.



We use the Fly algorithm, an artificial evolution strategy,
to reconstruct positron emission tomography (PET) images. The algorithm iteratively
optimises the position of 3D points. It eventually produces a point cloud, which needs
to be voxelised to produce volume data that can be used with conventional medical image
software. However, resulting voxel data is noisy. In our test case with 6,400 points
the normalised cross-correlation (NCC) between the reference and the reconstruction is
85.53%; with 25,600 points it is 93.60%. This paper introduces a more robust
3D voxelisation method based on implicit modelling using metaballs to overcome
this limitation. With metaballs, the NCC with 6,400 points increases up to 92.21%;
and up to 96.26% with 25,600 points.
|
|
[2]
|
 F. P. Vidal, J. Louchet, É. Lutton, and J.-M. Rocchisani.
PET reconstruction using a cooperative coevolution strategy in
LOR space.
In IEEE Nuclear Science Symposium Conference Record, pages
3363-3366, Orlando, Florida, October 2009. IEEE.
F. P. Vidal, J. Louchet, É. Lutton, and J.-M. Rocchisani.
PET reconstruction using a cooperative coevolution strategy in
LOR space.
In IEEE Nuclear Science Symposium Conference Record, pages
3363-3366, Orlando, Florida, October 2009. IEEE.


This paper presents preliminary results of a novel method that takes
advantage of artificial evolution for positron emission tomography
(PET) reconstruction. Fully 3D tomographic reconstruction in PET
requires high computing power and leads to many challenges. To date,
the use of such methods is still restricted due to the heavy computing
power needed. Evolutionary algorithms have proven to be efficient
optimisation techniques in various domains. However the use of evolutionary
computation in tomographic reconstruction has been largely overlooked.
We propose a computer-based algorithm for fully 3D reconstruction
in PET based on artificial evolution and evaluate its relevance.
Keywords: Positron emission tomography, genetic algorithms, optimization methods
|
 Cameron C. Gray, Shatha F. Al-Maliki, & Franck P. Vidal.
Data exploration in evolutionary reconstruction of PET images.
Genetic Programming and Evolvable Machines, 19(3):391-419,
September 2018.
Cameron C. Gray, Shatha F. Al-Maliki, & Franck P. Vidal.
Data exploration in evolutionary reconstruction of PET images.
Genetic Programming and Evolvable Machines, 19(3):391-419,
September 2018.



 Z. Ali Abbood, J. Lavauzelle, É Lutton, J.-M. Rocchisani, J. Louchet, and F. P. Vidal.
Voxelisation in the 3-D Fly Algorithm for PET.
Swarm and Evolutionary Computation, 36:91-105,
October 2017.
Z. Ali Abbood, J. Lavauzelle, É Lutton, J.-M. Rocchisani, J. Louchet, and F. P. Vidal.
Voxelisation in the 3-D Fly Algorithm for PET.
Swarm and Evolutionary Computation, 36:91-105,
October 2017.



 F. P. Vidal, P.-F. Villard, and É. Lutton.
Tuning of patient specific deformable models using an adaptive
evolutionary optimization strategy.
IEEE Transactions on Biomedical Engineering, 59(10):2942-2949,
October 2012.
F. P. Vidal, P.-F. Villard, and É. Lutton.
Tuning of patient specific deformable models using an adaptive
evolutionary optimization strategy.
IEEE Transactions on Biomedical Engineering, 59(10):2942-2949,
October 2012.




 Zainab Ali Abbood, and Franck P. Vidal. Basic, dual, adaptive, and directed mutation operators in the Fly algorithm.
In Biennial International Conference on Artificial Evolution, volume ???? of
Lecture Notes in Computer Science, pages ???-???, Paris, France,
October 2017. Springer, Heidelberg.
Zainab Ali Abbood, and Franck P. Vidal. Basic, dual, adaptive, and directed mutation operators in the Fly algorithm.
In Biennial International Conference on Artificial Evolution, volume ???? of
Lecture Notes in Computer Science, pages ???-???, Paris, France,
October 2017. Springer, Heidelberg.



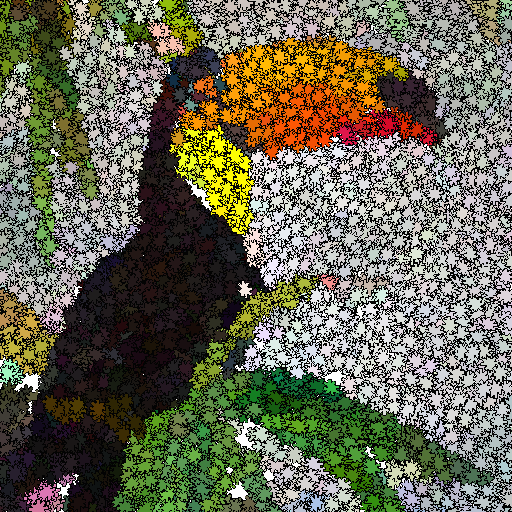 Z. Ali Abbood, O. Amlal, and F. P. Vidal.
Evolutionary Art Using the Fly Algorithm.
In Applications of Evolutionary Computation, volume 10199 of
Lecture Notes in Computer Science, pages 455-470, Amsterdam, The Netherlands,
April 2017. Springer, Heidelberg.
Z. Ali Abbood, O. Amlal, and F. P. Vidal.
Evolutionary Art Using the Fly Algorithm.
In Applications of Evolutionary Computation, volume 10199 of
Lecture Notes in Computer Science, pages 455-470, Amsterdam, The Netherlands,
April 2017. Springer, Heidelberg.







 P.-F. Villard, F. P. Vidal, F. Bello, and N. W. John.
A method to compute respiration parameters for patient-based
simulators.
In Proceeding of Medicine Meets Virtual Reality 19 - NextMed
(MMVR19), volume 173 of Studies in Health Technology and Informatics,
pages 529-533, Newport Beach, California, February 2012. IOS Press.
Winner of the best poster award.
P.-F. Villard, F. P. Vidal, F. Bello, and N. W. John.
A method to compute respiration parameters for patient-based
simulators.
In Proceeding of Medicine Meets Virtual Reality 19 - NextMed
(MMVR19), volume 173 of Studies in Health Technology and Informatics,
pages 529-533, Newport Beach, California, February 2012. IOS Press.
Winner of the best poster award.


 F. P. Vidal, É. Lutton, J. Louchet, and J.-M. Rocchisani.
Threshold selection, mitosis and dual mutation in cooperative
coevolution: application to medical 3D tomography.
In International Conference on Parallel Problem Solving From
Nature (PPSN'10), volume 6238 of Lecture Notes in Computer Science,
pages 414-423, Krakow, Poland, September 2010. Springer, Heidelberg.
F. P. Vidal, É. Lutton, J. Louchet, and J.-M. Rocchisani.
Threshold selection, mitosis and dual mutation in cooperative
coevolution: application to medical 3D tomography.
In International Conference on Parallel Problem Solving From
Nature (PPSN'10), volume 6238 of Lecture Notes in Computer Science,
pages 414-423, Krakow, Poland, September 2010. Springer, Heidelberg.



 F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
New genetic operators in the Fly algorithm: application to medical
PET image reconstruction.
In Applications of Evolutionary Computation, volume 6024 of
Lecture Notes in Computer Science, pages 292-301, Istanbul, Turkey,
April 2010. Springer, Heidelberg.
Nominated for best paper award.
F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
New genetic operators in the Fly algorithm: application to medical
PET image reconstruction.
In Applications of Evolutionary Computation, volume 6024 of
Lecture Notes in Computer Science, pages 292-301, Istanbul, Turkey,
April 2010. Springer, Heidelberg.
Nominated for best paper award.



 F. P. Vidal, D. Lazaro-Ponthus, S. Legoupil, J. Louchet, É. Lutton, and
J.-M. Rocchisani.
Artificial evolution for 3D PET reconstruction.
In Proceedings of the 9th international conference on Artificial
Evolution (EA'09), volume 5975 of Lecture Notes in Computer Science,
pages 37-48, Strasbourg, France, October 2009. Springer, Heidelberg.
F. P. Vidal, D. Lazaro-Ponthus, S. Legoupil, J. Louchet, É. Lutton, and
J.-M. Rocchisani.
Artificial evolution for 3D PET reconstruction.
In Proceedings of the 9th international conference on Artificial
Evolution (EA'09), volume 5975 of Lecture Notes in Computer Science,
pages 37-48, Strasbourg, France, October 2009. Springer, Heidelberg.



 F. P. Vidal, M. Folkerts, N. Freud, and S. Jiang.
GPU accelerated DRR computation with scatter.
Medical Physics, 38(6):3455-3456, July 2011.
F. P. Vidal, M. Folkerts, N. Freud, and S. Jiang.
GPU accelerated DRR computation with scatter.
Medical Physics, 38(6):3455-3456, July 2011.



 F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
Flies for PET: An artificial evolution strategy for image
reconstruction in nuclear medicine.
Medical Physics, 37(6):3139, July 2010.
F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
Flies for PET: An artificial evolution strategy for image
reconstruction in nuclear medicine.
Medical Physics, 37(6):3139, July 2010.



 Zainab Ali Abbood, Jean-Marie Rocchisani, and Franck P. Vidal.
Visualisation of PET data in the Fly Algorithm.
In Katja Bühler, Lars Linsen, and Nigel W. John, editors,
Eurographics Workshop on Visual Computing for Biology and Medicine, pages
211--212. The Eurographics Association, 2015.
Zainab Ali Abbood, Jean-Marie Rocchisani, and Franck P. Vidal.
Visualisation of PET data in the Fly Algorithm.
In Katja Bühler, Lars Linsen, and Nigel W. John, editors,
Eurographics Workshop on Visual Computing for Biology and Medicine, pages
211--212. The Eurographics Association, 2015.



 F. P. Vidal, J. Louchet, É. Lutton, and J.-M. Rocchisani.
PET reconstruction using a cooperative coevolution strategy in
LOR space.
In IEEE Nuclear Science Symposium Conference Record, pages
3363-3366, Orlando, Florida, October 2009. IEEE.
F. P. Vidal, J. Louchet, É. Lutton, and J.-M. Rocchisani.
PET reconstruction using a cooperative coevolution strategy in
LOR space.
In IEEE Nuclear Science Symposium Conference Record, pages
3363-3366, Orlando, Florida, October 2009. IEEE.




![Validate my Atom 1.0 feed [Valid Atom 1.0]](img/valid-atom.png)